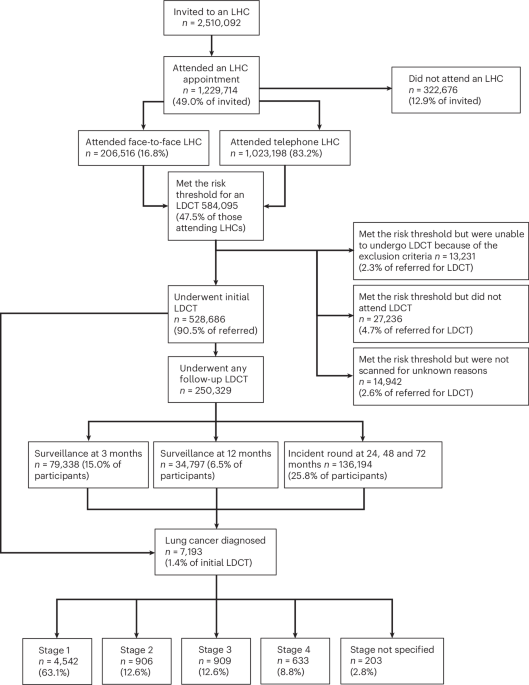
Study population
In this study, we analyzed data to examine the associations among accelerated biological aging, lung cancer risk, and all-cause mortality among individuals with lung cancer. Data were drawn from the U.S. National Health and Nutrition Examination Survey (NHANES), a continuous cross-sectional study conducted over 2-year cycles from 2001 to 2020. We included participants aged > 18 years with available data on the clinical biomarkers required for the calculation of PhenoAge and lung cancer status on the basis of self-reported physician diagnoses. For the mortality analysis, we further excluded individuals with missing death records.
This study was conducted in accordance with the Declaration of Helsinki and approved by the National Center for Health Statistics (NCHS) Research Ethics Review Board19. All participants provided written informed consent.
Ascertainment of outcomes
In the NHANES, all participants completed a questionnaire that included a cancer-related question: “Have you ever been told by a doctor or other health professional that you had cancer or a malignancy of any kind?” The participants who responded “yes” were asked a follow-up question (“What kind of cancer was it?“) to identify the participants who were diagnosed with lung cancer.
For the longitudinal analysis, mortality status was ascertained from the NHANES Public Use Linked Mortality File, which provides vital status follow-up data through record linkage, enabling survival analyses of NHANES participants20.
Procedures
PhenoAge was computed for all NHANES participants as a measure of biological aging which captures the influence of clinical biomarkers on health outcomes and aligns with previous work demonstrating its ability to predict disease risk and aging-related outcomes more effectively than chronological age13. PhenoAge was derived using the Gompertz proportional hazard model, which is widely recognized in aging research for modeling mortality risk and its exponential increase with age. This model is particularly well-suited for aging studies as it reflects the acceleration of aging-related mortality risk over time. The calculation of PhenoAge follows a well-established protocol, where two Gompertz proportional hazard models are parameterized: The first model incorporates ten biomarkers and chronological age to estimate an individual’s biological aging status. The second model uses chronological age alone as the predictor variable, serving as a baseline comparison to evaluate whether biological aging (PhenoAge) provides additional predictive value beyond chronological age alone (Levine, 2013). In accordance with the protocol previously outlined in related studies, the NHANES III dataset was employed as the reference population, with an individual’s PhenoAge reflecting the chronological age of an individual with the same mortality risk21. The following formula was employed for the purpose of determining the PhenoAge.
$$\beginaligned \textPhenoAge = & 143.5671 \\ & + \frac\ln [ – 0.0059383581 \times \ln [1 – \text mortality risk ]]0.08548908 \\ & \textmortality\;\text risk = 1 – e^{{ – e^x\frac\exp (120x\gamma ) – 1\gamma }} \\ & \gamma = 0.007354285 \\ \endaligned$$
where xb = −19.907−0.0336*Albumin + 0.0095*Creatinine + 0.1953 × Glucose-0.0120 × Lymphocyte Percent + 0.0268 × Mean Cell Volume + 0.3306 × Red Cell Distribution Width + 0.00188 × Alkaline Phosphatase + 0.0554 × White Blood Cell Count + 0.0804 × Chronological Age.
The concept of biological aging is defined as the value of biological age that exceeds an individual’s chronological age. Accelerated biological aging was determined through a calculation of the difference between the estimated biological age and the actual chronological age21.
In this study, based on their clinical relevance and existing literature12,22, the following variables were included: age, sex (male, female), race (non-Hispanic white, non-Hispanic black, Mexican American, other), education level (grade school or less, high school, more than high school), poverty status (ratio of family income to poverty), smoking status (never, former, current smoker), drinking status (never, former, light or moderate, heavy), and BMI. Additionally, history of emphysema was included in this study. The participants were classified according to their smoking status, which was divided into three categories: current smokers, former smokers, and nonsmokers. The term ‘current smoker’ was used to describe individuals who had smoked on a regular basis and had smoked at least 100 cigarettes in their lifetime. Former smokers had smoked at least 100 cigarettes and subsequently ceased this practice. Those who had never smoked were classified as nonsmokers, as were those who had smoked fewer than 100 cigarettes. The drinking status of the participants was categorized according to the following definitions: never (had consumed fewer than 12 alcoholic beverages in their lifetime), former (had consumed 12 or more alcoholic beverages in a single year but had not drunk any alcohol in the preceding year or had not drunk any alcohol in the preceding year but had consumed 12 or more alcoholic beverages at some point in their lifetime), light/moderate drinker (≤ 1 alcoholic beverage per day for women or ≤ 2 alcoholic beverages per day for men on average over the past year), or current heavier drinker (> 1 alcoholic beverage per day for women or > 2 alcoholic beverages per day for men on average over the past year). Individuals can be classified according to their body mass index (BMI) into one of three categories: ≤25, > 25 to < 30, or ≥ 30 kg/m2. The medical history of the participants regarding emphysema was established on the basis of the self-reported information provided by the participants.
Study design
In this study, we employed a two-stage design to examine the relationships among accelerated biological aging, lung cancer risk, and all-cause mortality among individuals with lung cancer, utilizing data from the NHANES. In Stage 1, we performed a cross-sectional analysis to investigate the association between PhenoAge acceleration and the prevalence of lung cancer at baseline. Lung cancer status was determined on the basis of self-reported diagnoses collected through NHANES questionnaires. In Stage 2, a longitudinal analysis was conducted to evaluate the association between PhenoAge acceleration and all-cause mortality among participants diagnosed with lung cancer. Mortality data were obtained from the NHANES Public Use Linked Mortality File, enabling survival analysis during the follow-up period.
Statistical analysis
In the present study, we employed NHANES data to ascertain the baseline characteristics of patients with and without lung cancer. The data are expressed as the means ± SDs (x ± s). The statistical analysis of the differences among the groups was conducted using either Student’s t test or one-way analysis of variance (ANOVA). For the purpose of multiple comparisons, the SNK or LSD method was employed. Qualitative data are presented as percentages and were analyzed via either a chi-square (x2) test or Fisher’s exact test, as appropriate. Missing data were handled by treating missing values for categorical variables (BMI, smoking status, drinking status, education level, and history of emphysema) as separate categories to preserve the full sample size and avoid potential selection bias. The reported P value was two-sided, and a value of less than 0.05 was considered to indicate statistical significance. In alignment with the NHANES analytic guidelines, the present analyses accounted for the complex sampling design, including sampling weights, stratification (strata), and clustering (PSU, primary sampling units). All statistical models incorporated NHANES survey design variables to ensure correct variance estimation and population inference. Furthermore, all analyses were adjusted for potential confounding variables.
In this study, PhenoAge acceleration (PhenoAgeAccel) was initially calculated as a continuous variable using the R package BioAge. To explore the distribution of PhenoAgeAccel across participants, we also categorized it into quartiles on the basis of the PhenoAgeAccel distribution in the study cohort, with cutoffs corresponding to the 25th, 50th (median), and 75th percentiles. Furthermore, PhenoAge acceleration was computed by regressing PhenoAge against chronological age at the time of biomarker measurement, with the residuals defining the PhenoAgeAccel.
In logistic regression models, PhenoAge acceleration was assessed both as a continuous variable and as a binary variable. ‘Accelerated biological aging’ was defined as a PhenoAge acceleration value exceeding zero, whereas ‘nonaccelerated aging’ was defined as a PhenoAge acceleration value of zero or less. Given the presence of zero inflation (many participants exhibited no evidence of accelerated aging), we used both continuous and binary classifications of PhenoAge acceleration to assess its impact on lung cancer risk.
The clinical outcome of all-cause mortality over time was described with Kaplan–Meier survival curves and compared by the log-rank test. Furthermore, Cox proportional hazards regression models were used to evaluate the relationships between PhenoAge and mortality in individuals with lung cancer. The confounders were selected on the basis of clinical interest, previous scientific literature and the identification of all significant covariates in the univariate analysis. The objective was to evaluate the association between lung cancer incidence and PhenoAge acceleration, and for this purpose, several models were employed. The crude model was not adjusted for any variables, whereas Model 1 was adjusted for age, sex, and race. Model 2 was further adjusted for education level, the family income-to-poverty ratio, smoking status, drinking status, BMI, and history of emphysema. This stepwise adjustment approach is widely used in epidemiological studies to evaluate whether biological aging provides additional predictive value beyond demographic and lifestyle factors22. It allows for a clearer understanding of the independent association between PhenoAge acceleration and lung cancer risk. The P value for the trend across increasing exposure groups was calculated using integer values (1, 2, 3, and 4). A restricted cubic spline (RCS) regression model was employed to assess the dose‒response relationships between biological age (PhenoAge) and the risk of developing lung cancer, as well as between biological age and all-cause mortality in individuals with lung cancer. The model was fitted using three knots placed at the 25th, 50th, and 75th percentiles of the PhenoAge distribution to capture the nonlinear relationships and better understand how biological age influences both outcomes.
We performed several sensitivity analyses. First, participants who had values that fell within the 1% extremes of the PhenoAge acceleration range were excluded to rule out the effects of extreme values. Second, the associations between PhenoAge acceleration and lung cancer and all-cause mortality were reanalyzed without the consideration of complex sampling designs. Third, to determine the impact of various factors on the relationship between lung cancer and biological aging, we conducted a subgroup analysis and an interactive test. Fourth, we conducted sensitivity analyses to assess whether missingness influenced the study outcomes. Finally, we applied several additional association inference models: propensity score adjustment (PSA), propensity score matching (PSM)23, inverse probability of treatment weighting (IPTW)24, standardized mortality ratio weighting (SMRW)25, pairwise algorithm (PA)26, and overlap weight (OW)27. The calculated effect sizes and P values from all these models were reported and compared. All the analyses were performed with the statistical software packages R3.3.2 ( The R Foundation) and the Free Statistics analysis platform (Version 1.9, Beijing, China,
link