Study population
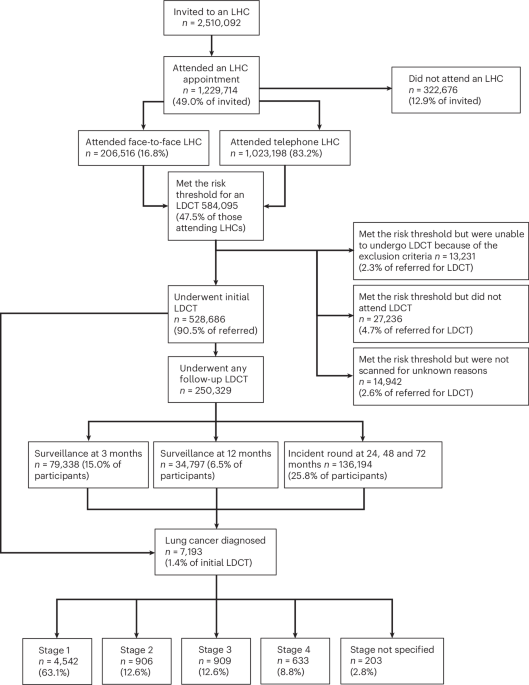
This study is based on data from the SEPAGES French parent–child cohort. The population of the study lives within 80 km around the center of Grenoble in the Alpes, France. Briefly, 484 pregnant women, with pregnancy duration less than 19 weeks and with a singleton pregnancy, older than 18 years old and affiliated to the French national security system, were recruited between 2015 and 201721. Their partner and children were also recruited. Sociodemographic and health data were collected using a combination of questionnaires, interviews, and clinical examinations during and after pregnancy. Exposure information were collected using personal samplers and immunological information were collected using blood samples. The present analysis is based on 270 mothers with exposure assessed to at least one of the measured air pollutants during pregnancy and immunological measurements at the end of the exposure assessment week (Fig. 1). To be included in the analysis, mothers had to have the blood samples collected within 2 days after the pollutant sampling period to assess the immune system’s status in relation to the exposure measurements. Participants signed an informed consent and the study protocol received approval from the French data privacy institution (Commission Nationale de l’Informatique et des Libertés, CNIL—n°914138) and the Comité de Protection des Personnes Sud-Est V (CPP—2013-A01491-44). All methods were performed in accordance with the relevant guidelines and regulations.

Flow chart for the selection of the study population. Abbreviations: PM2.5 particulate matter with diameter ≤ 2.5 μm; NO2 nitrogen dioxide; OP: oxidative potential; PHA phytohemagglutinin; R848 resiquimod.
Personal exposure assessment to air pollutants
Women in the SEPAGES cohort wore or kept nearby personal air samplers placed in a wearable backpack to measure their personal exposures to PM2.5 (MicroPEM™ active air sampler; RTI International, USA) and NO2 (Passam AG passive air sampler; Switzerland) for 7 (13%) to 8 (87%) consecutive days22,23,24. The measurement week took place during the second (81%) or the third trimester (19%) of pregnancy (median [min–max] gestational age = 19 weeks14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36).
NO2 concentration was measured using spectrophotometry following established methods25. PM2.5 filters from the MicroPEM were weighed before and after sampling at RTI International (USA), using a microbalance (Mettler Toledo UMX2) placed in an environmental chamber maintained at a temperature of 21 °C and 35% relative humidity. PM2.5 mass concentration was calculated by dividing PM mass collected during the week and measured by gravimetric analysis, by the air volume sampled during the measurement week (µg/m3). PM filters were cold-stored until OP analysis. The protocol for OP measurement was previously published22,24, based on the protocol established by Calas et al.26,27. Briefly, PM2.5 were extracted from filters into a simulated interstitial lung fluid consisting in a mixture of 1,2-dipalmitoylphosphatidylcholine (DPPC), to reach a final concentration of 10 µg/mL. Extracts were incubated at 37 °C for 75 min under vortex agitation prior to analysis using the dithiothreitol (DTT) and the ascorbic acid (AA) assays. A 96-well plate (CELLSTRAR, Greiner-Bio) was used to mix the extracts with DTT or AA solutions. For the AA assay, the absorbance at 265 nm (TECAN spectrophotometer Infinite M200 Pro) is measured over time to evaluate AA consumption by PM2.5 extract, for a total reaction time of 30 min. For the DTT assay, the absorbance at 412 nm measured the formation of the 2-nitro-5-thiobenzoic acid (TNB), which is the reaction product of the remaining DTT and dithionitrobenzoic acid (DTNB), for a total reaction time of 30 min. Samples were analyzed in triplicates, and the mean was calculated for each sample. For both assays, consumption rates were then normalized by the mass of PM of the extract (OPm, in nmol/min/µg), or by the corresponding air volume sampled (OPv, nmol/min/m3).
Maternal immune function
Blood samples were collected by trained field workers, within a maximum of 48 h after the end of the exposure measurement week, following the procedure published by Manches et al.20. Briefly, blood was collected in BD Medical 368886 vacutainer tubes (lithium heparin) for immunological analyses (cell culture and plasma separation), and in BD Medical 368861 vacutainer tubes (EDTA) for cell counting. They were transported to the Etablissement Français du Sang (EFS) in coolers, placed on a rotating device for at least 5 min to ensure homogeneous cell content, and were then processed within 24 h after collection.
Innate and adaptative immunity of the women were measured at baseline and after a 24-h ex vivo activation of whole blood at 37 °C using dendritic cell activator Resiquimod (R848, InvivoGen, 5 μg/mL) and T cell activator phytohaemagglutinin (PHA, Oxoid, 10 μg/mL), as previously described by Manches et al.20.
Briefly, cytokines were measured in the culture supernatant (for activated cells) or in plasma (for baseline cytokines) by cytometric bead arrays (BD™ CBA Human cytokines Flex Set that is a bead-based immunoassay capable of simultaneously measuring several cytokines in biological fluids, BD Biosciences).
Among the 29 cytokines that were measured (12 at baseline, 9 after PHA-activation and 8 after R848 activation) only those with at least 70% of detected values were considered20. Hence, for the samples activated with PHA, the overall activity of T lymphocytes (T helpers Th1, Th2, Th9, Th17, and regulatory Treg) was assessed by quantifying the levels of IL-2, TNF-α, interferon (IFN) γ, IL-13, IL-17a, IL-9 and IL-10. For the samples activated with R848, the overall activity of dendritic cells was evaluated by quantifying TNF-α, IL-10, IL-6, IL-8, IFN-α, IFN-γ, IL-1β, and IL-12p70. For the non-activated sample, the basal state of the immune system was quantified by IL-8, monocyte chemoattractant protein-1 (MCP1) and regulated on activation, normal T cell expressed and secreted (RANTES). The concentrations below the limit of detection were imputed by a fill-in approach, that randomly selects values between 0 and the LOD based on the underlying distribution28,29. Due to their skewed distribution, cytokine concentrations were log10 transformed.
Since between-participant technical variability related to the experimentation can lead to measurement error, a two-step standardization method based on regression residuals30 was used to correct, when necessary, cytokine concentrations. The same standardized variables as previously described by Manches et al.20 were used. Briefly, the technical variables considered were: (1) for baseline cytokines: analytical batch, time between sample collection and reception, time between sample reception and analysis; (2) for activated cytokines the same variables were used, together with the duration of the activation, R848 or PHA age at the time of sample activation, and storage duration.
Statistical analysis
Summary statistics (mean [sd] or median [Q1-Q3]) were calculated for air pollutant exposure assessments, cytokine levels, and covariates. A correlation matrix (Pearson’s r) was calculated between the cytokine levels and between the air pollutant concentrations. Univariate and adjusted linear regressions were conducted to estimate the associations between each air pollutant exposure and each cytokine level. Each exposure and log-transformed cytokine variable were divided by the interquartile range (IQR), to facilitate comparison of the beta estimates. Potential confounders were selected from the existing literature31,32,33, and using a directed acyclic graph (see Supplementary Fig. S1 online) and included: age of the women (continuous), BMI before pregnancy (continuous), active or passive smoking (active smoking in the 12 months prior to pregnancy, or active or passive smoking during pregnancy; binary: yes/no), educational level (binary: < master’s degree, ≥ master’s degree), leukocyte count (continuous), gestational age at sampling (continuous), and sampling season (4 categories with winter corresponding to January-March, spring to April-June, summer to July–September, and fall to October-December). To avoid reduction of the sample size due to missing data in cofactors (20 individuals had missing value for at least one covariate), multiple imputations (n = 20 datasets) were performed using Multivariate Imputation by Chained Equations (package mice, R).
In addition, sensitivity analyses were carried out to assess the robustness of the results to: (1) extreme values (after exclusion of 1% lowest and 1% highest exposure and cytokine concentrations), (2) influential values (after exclusion of values with a Cook’s distance exceeding 4/n, with n being the number of participants in the main analysis), (3) the set of confounders, with models excluding the leucocyte counts among cofactors and models including history of asthma and rhinitis which could lie in the causal path between air pollution and cytokine levels, and (4) considering bi-pollutant models, e.g. adjusting for NO2 in the PM2.5 and OP models, and adjusting for PM2.5 or OP in the NO2 models. Following statistical recommendations, interpretation of statistical tests was based on examining effects’ magnitude and their 95% confidence intervals and precise p-values (not whether p-values are above or below 0.05)34,35. All analyses were conducted using the statistical software R (version 4.2).
link