We conducted this study using data from the global Collaborative Network within the TriNetX Research Network, which comprises real-time, de-identified electronic health records from 146 healthcare organizations, including hospitals, primary care providers, and specialty clinics. Available data included demographics, diagnoses (ICD-10-CM), procedures (ICD-10-PCS and CPT), medications, laboratory tests (LOINC), and healthcare utilization.
TriNetX is a global federated research platform widely used for large-scale observational studies. For this analysis, we identified a pediatric cohort between July 1, 2016, and June 4, 2025, to evaluate the prognostic value of serum cystatin C in predicting progression to chronic kidney disease among children with congenital urologic abnormalities. Given the anonymized nature of the dataset, informed consent was not required.
Ethics statement
Regarding ethical approval, per the TriNetX publication guidelines, “This retrospective study is exempt from informed consent. The data reviewed is a secondary analysis of existing data, does not involve intervention or interaction with human subjects, and is de-identified per the de-identification standard defined in Section §164.514(a) of the HIPAA Privacy Rule. The process by which the data is de-identified is attested to through a formal determination by a qualified expert as defined in Section §164.514(b)(1) of the HIPAA Privacy Rule. This formal determination by a qualified expert was refreshed in December 2020.” ( accessed January 12, 2025).
Cohort construction and patient selection
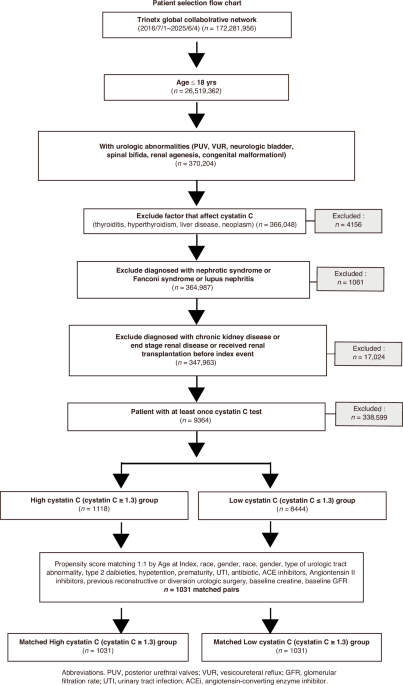
A flowchart of the cohort construction from 168,953,527 participants enrolled in the TriNetX Global Collaborative Network between 1 July 2016 and 4 June 2025 is provided in Fig. 1 and Supplementary Fig. S1. Pediatric patients aged 18 years or younger were included in the cohort. Patients with urologic abnormalities, including posterior urethral valves (PUV), vesicoureteral reflux (VUR), neurologic bladder, spina bifida, renal agenesis, multicystic dysplastic kidney, ureteropelvic junction obstruction, ureterovesical junction obstruction, and cloaca malformation were included. Patients diagnosed with neoplasm, nephrotic syndrome, lupus nephritis, Fanconi syndrome, or those who received a diagnosis of chronic kidney disease, end-stage renal disease, or received renal transplantation or dialysis before the index event were excluded. Additionally, patients without cystatin C testing and those with factors affecting cystatin C levels (thyroiditis, hyperthyroidism, liver disease, neoplasms) were excluded. After exclusion, the study population (n = 9562) was divided into high cystatin C (n = 1118) and low cystatin C (n = 8444) groups based on a threshold of 1.3 mg/L, which reflect the Kidney Disease: Improving Global Outcomes (KDIGO) guideline recommendation to incorporate cystatin C for improved CKD detection in vulnerable populations, and with published pediatric reference thresholds for identifying chronic kidney disease (CKD).11,12,13,14

Flow diagram illustrating patient identification, inclusion and exclusion criteria, propensity score matching, and final cohort formation from the TriNetX database. PUV posterior urethral valves, VUR vesicoureteral reflux, GFR glomerular filtration rate, UTI urinary tract infection, ACEi angiotensin-converting enzyme inhibitor.
In our cohort, 1:1 propensity score matching was performed based on age at index, race, sex, type of urologic tract abnormality, urinary tract infection, type 2 diabetes, hypertension, antibiotic use, steroid use, and urologic surgery, including posterior urethral valve ablation, pyeloplasty, ureteroscope dilatation, or internal stent drainage. Additionally, baseline serum creatinine, estimated glomerular filtration rate (eGFR), and albumin/creatine ratio were included in the matching process. After matching, a total of 1031 patients with elevated serum cystatin C levels and 1031 matched controls with lower cystatin C levels were identified. These patients were longitudinally followed to evaluate renal outcomes and related complications. A complete list of ICD-10-CM codes used for cohort identification is provided in Appendix Table 1.
Index event
The index event was defined as the diagnosis of a urologic malformation followed by a measurement of serum cystatin C. Eligible index events could have occurred up to 10 years prior to the date of data extraction.
Pre-specified outcomes
The primary outcome was the incidence of major adverse kidney events (MAKE) occurring between 1 day after the index event and up to 10 years of follow-up. MAKE was defined as a composite of the following:
-
1.
Progression to CKD: Diagnosis of Chronic kidney disease (ICD-10-CM: N18) or End stage renal disease (ICD-10-CM: N18.6)
-
2.
Need for dialysis:
-
CPT:90937 Hemodialysis procedure requiring repeated evaluation(s) with or without substantial revision of dialysis prescription
-
CPT:90935 Hemodialysis procedure with single evaluation by a physician or other qualified health care professional
-
CPT:90947 Dialysis procedure other than hemodialysis (e.g., peritoneal dialysis, hemofiltration, or other continuous renal replacement therapies) requiring repeated evaluations by a physician or other qualified health care professional, with or without substantial revision of dialysis prescription
-
CPT:90945 Dialysis procedure other than hemodialysis (e.g, peritoneal dialysis, hemofiltration, or other continuous renal replacement therapies), with single evaluation by a physician or other qualified health care professional
-
CPT 1012752 Hemodialysis procedure
-
CPT 1012740 Dialysis services and procedures
-
ICD-10-CM Z99.2 dependence on renal dialysis
-
SNOMED peritoneal dialysis
-
SNOMED 108241001 Dialysis procedure
-
SNOMED chronic peritoneal dialysis
-
SNOMED continuous hemodialysis
-
SNOMED maintenance hemodialysis
-
SNOMED 238318009 continuous ambulatory peritoneal dialysis
-
-
3.
Need for kidney transplantation
-
ICD10CM:Z94.0 Kidney transplant status
-
ICD10PCS:0TY00Z0 Transplantation of Right Kidney, Allogeneic, Open Approach
-
ICD10PCS:0TY10Z0 Transplantation of Left Kidney, Allogeneic, Open Approach
-
ICD10PCS:0TY00Z1 Transplantation of Right Kidney, Syngeneic, Open Approach
-
ICD10PCS:0TY10Z1 Transplantation of Left Kidney, Syngeneic, Open Approach
-
CPT:1008104: Backbench reconstruction of cadaver or living donor renal allograft prior to transplantation
-
CPT:50323 Backbench standard preparation of cadaver donor renal allograft prior to transplantation, including dissection and removal of perinephric fat, diaphragmatic and retroperitoneal attachments, excision of adrenal gland, and preparation of ureter(s), renal vein(s), and renal artery(s), ligating branches, as necessary
-
CPT:50327 Backbench reconstruction of cadaver or living donor renal allograft prior to transplantation; venous anastomosis, each
-
CPT:50325 Backbench standard preparation of living donor renal allograft (open or laparoscopic) prior to transplantation, including dissection and removal of perinephric fat and preparation of ureter(s), renal vein(s), and renal artery(s), ligating branches, as necessary (UMLS: CPT:50325)
-
CPT:50328 Backbench reconstruction of cadaver or living donor renal allograft prior to transplantation; arterial anastomosis, each
-
CPT:1008098 Renal Transplantation Procedures
-
-
4.
eGFR ≤60 mL/min/1.73 m² based on CKD-EPI formula (LOINC: 62238-1), MDRD formula (TNX:8001), Schwartz formula (LNC:50384-7)
-
5.
Albuminuria
-
Albumin/Creatinine ratio in 24-hour urine ≥3 mg/mmol (LOINC: 76401-9)
-
Albumin/Creatinine mass ratio in 24-hour urine ≥30 mg/g (LOINC: 13705-9)
-
Albumin/Creatinine [Ratio] in Urine (LOINC:32294-1) ≥ 3 mg/mmol
-
Albumin/Creatinine [Mass Ratio] in 24 hour Urine ≥30 mg/g (TNX: LG34557-5)
-
Albumin/Creatinine [Mass Ratio] in Urine ≥30 mg/g (LOINC: 9318-7)
-
With any MAKE occurrence within 1 month prior to the index event were excluded. The secondary outcomes included each individual component of the composite.
Covariates
To adjust for differences in baseline characteristics between the two groups, we included several covariates in the analysis. Demographic variables such as age, sex, and race were incorporated, along with comorbidities associated with impaired renal function, including congenital urologic malformations (ICD-10: Q60–Q64), congenital hydronephrosis (Q62), vesicoureteral reflux (VUR; Q62.7), cystic kidney disease(Q61), Congenital posterior urethral valves (Q64.2) urinary tract infection (N39), hypertension (I10–I1A), and type 2 diabetes (E11). We also accounted for exposures that may influence renal function or serum cystatin C levels, including the use of antibiotics, non-steroidal anti-inflammatory analgesics, angiotensin-converting enzyme inhibitors, angiotensin II receptor blockers, and corticosteroids, as well as prior urologic surgery. Laboratory parameters considered as potential confounders included baseline estimated glomerular filtration rate (eGFR), serum creatinine, and urine albumin-to-creatinine ratio. A complete list of ICD-10-CM codes, Current Procedural Terminology (CPT) codes, Anatomical Therapeutic Chemical (ATC) classifications, and Veterans Affairs (VA) drug class codes used in the covariate analysis is provided in Appendix Table 2.
Statistical analyses
To minimize confounding, we employed propensity score matching (PSM) to create comparable groups with similar baseline characteristics. Using the built-in TriNetX function, we performed 1:1 greedy nearest neighbor matching based on age at index, sex, race, and key confounders such as prematurity, urologic abnormalities, urinary tract infection, and vesicoureteral reflux (VUR). Standardized differences (Std diff) were used to assess covariate balance between matched groups, with a value < 0.1 indicating acceptable balance.
To avoid reverse causality, follow-up started one day after the index event and continued for up to ten years. Hazard ratios (HRs) for progression to major adverse kidney events (MAKE) were estimated for patients with high versus low cystatin C levels. The proportional hazards assumption was tested using the generalized Schoenfeld residual method available in the TriNetX platform. If this assumption was violated, time-dependent HRs were calculated for different follow-up intervals.
All statistical analyses were conducted with a significance threshold of p < 0.05, and 95% confidence intervals (CI) were reported. The Kaplan–Meier method was used to estimate survival probabilities. Additionally, subgroup analyses were performed to evaluate the risk associated with each individual component of the composite MAKE outcome.
link